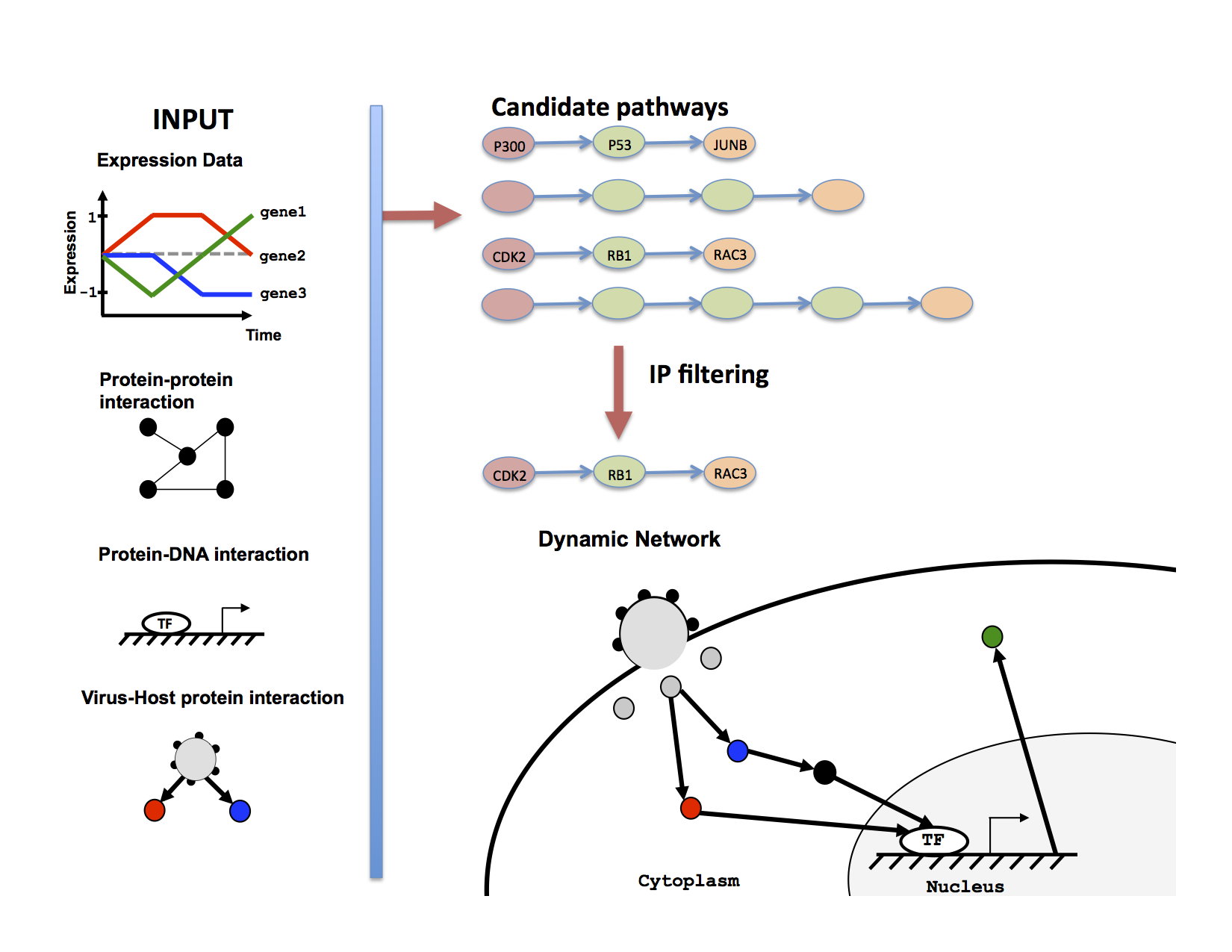
Most methods for reconstructing signaling and regulatory response
networks from high throughput data generate static models which cannot
distinguish between early and late stages in the response. Here we
present TimePath, a new method that integrates time series and static
datasets to reconstruct dynamic
models of host immune response. TimePath works by selecting a
subset of pathways that, together, explain the observed
dynamic responses using an Integer Programming formulation. Applying
TimePath to study human response to HIV-1 led to accurate
reconstruction of several known signaling and regulatory pathways and to
the identification of additional novel proteins. Each of these pathways
and proteins were assigned by TimePath to a specific temporal phase in
the response. We experimentally validated many of these assignments
shedding new light on the function of specific proteins and
demonstrating that treatments beyond the phase assigned by TimePath are
much less effective in reducing viral loads.
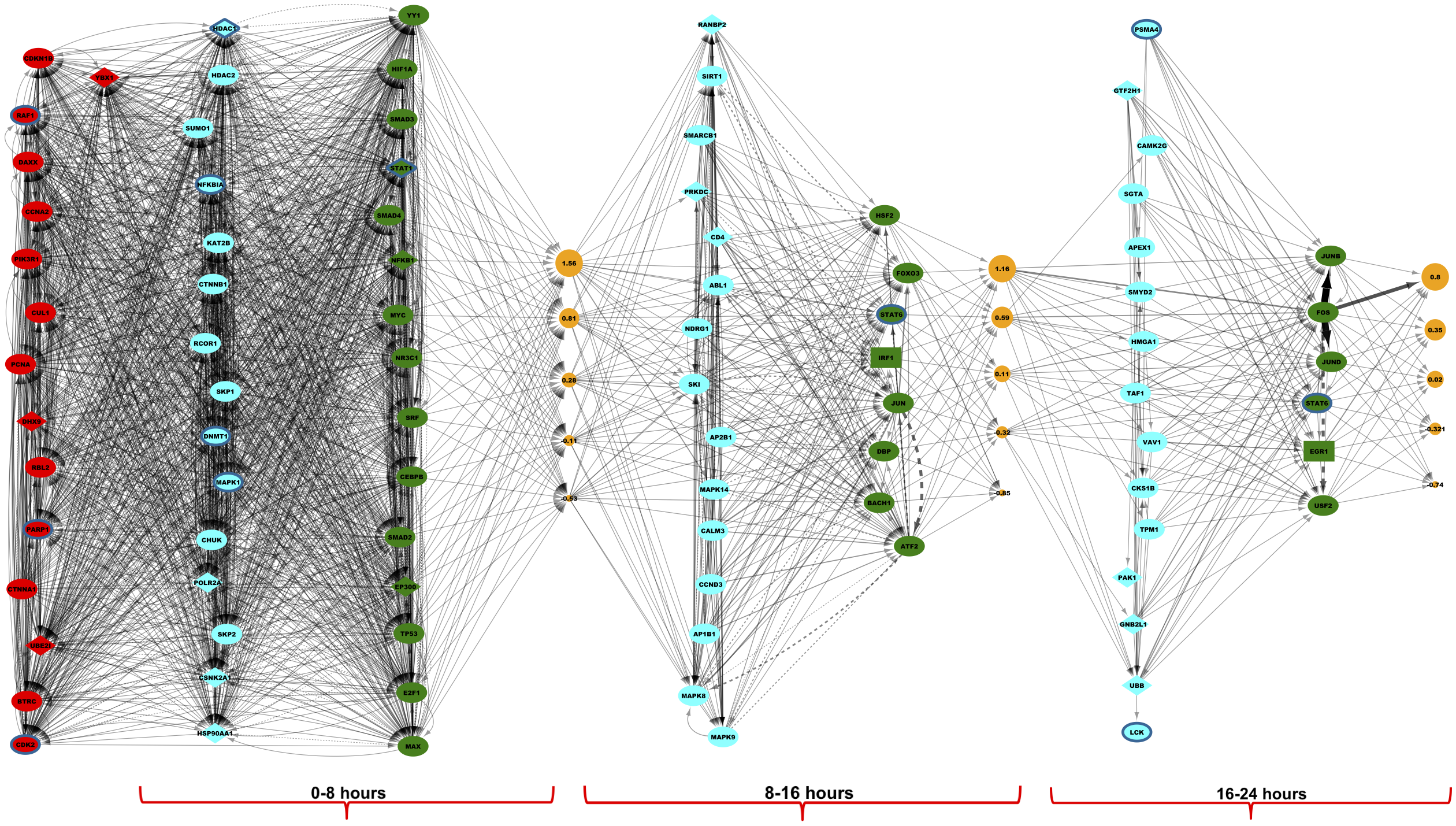
The figure above shows the method overview (left) and temporal signaling network (right) that TimePath learns for the response of the cell to infection by the HIV-1 virus.